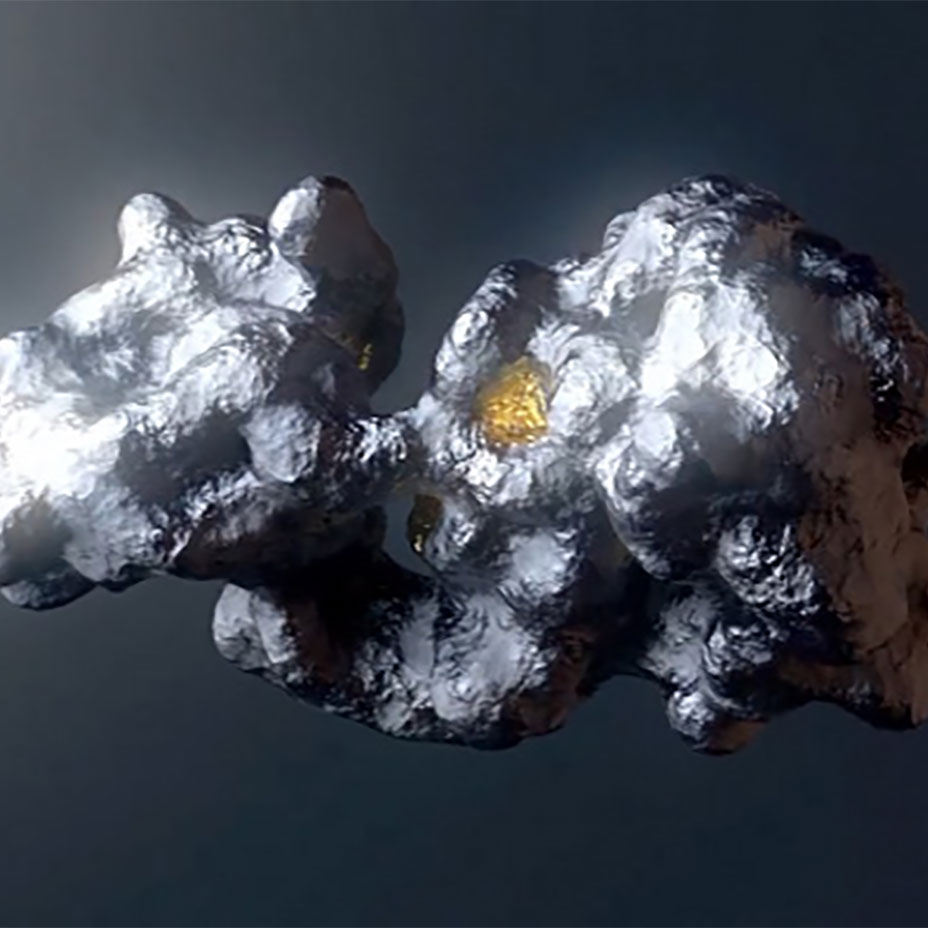
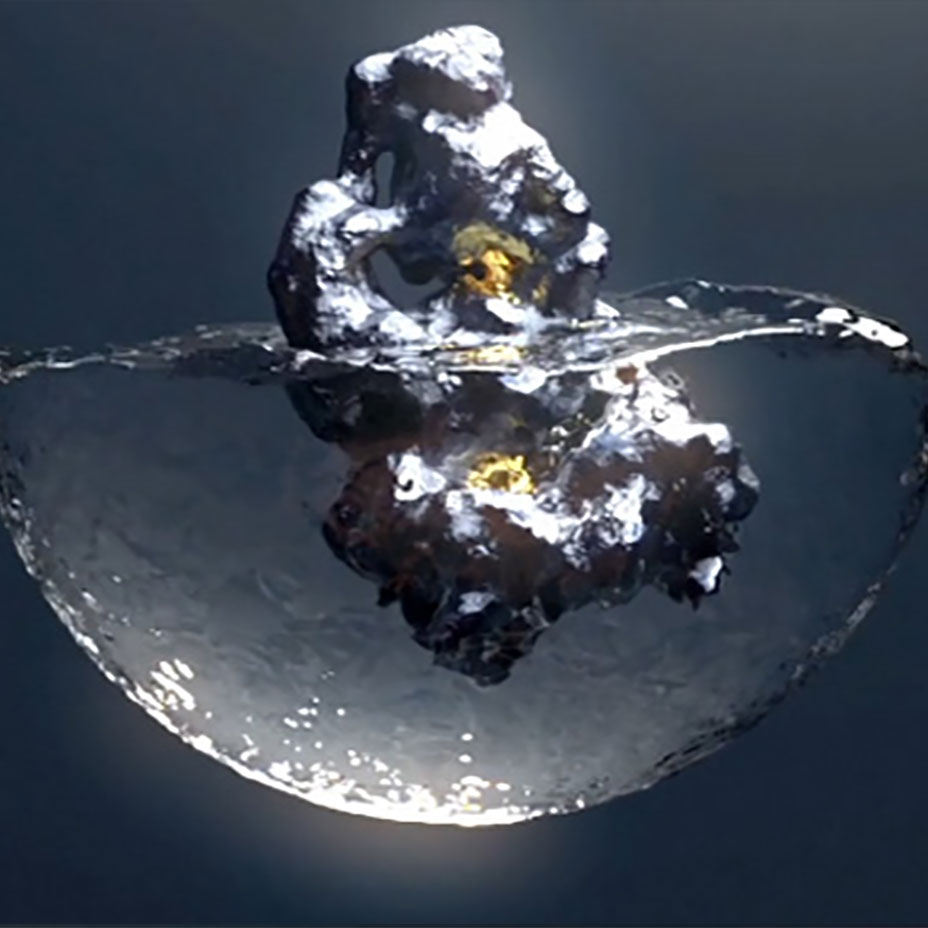
Our ReSOLVE™ drug discovery platform integrates our structural biology and protein science expertise with our proprietary computational chemistry tools to address current industry limitations. ReSOLVE™ is the only platform that can: model all conformations of a protein, identify druggable pockets, understand the precise and dynamic water networks of a pocket, and use those water networks to generate a blueprint for small molecule therapeutics, which we refer to as a hydrocophore™.
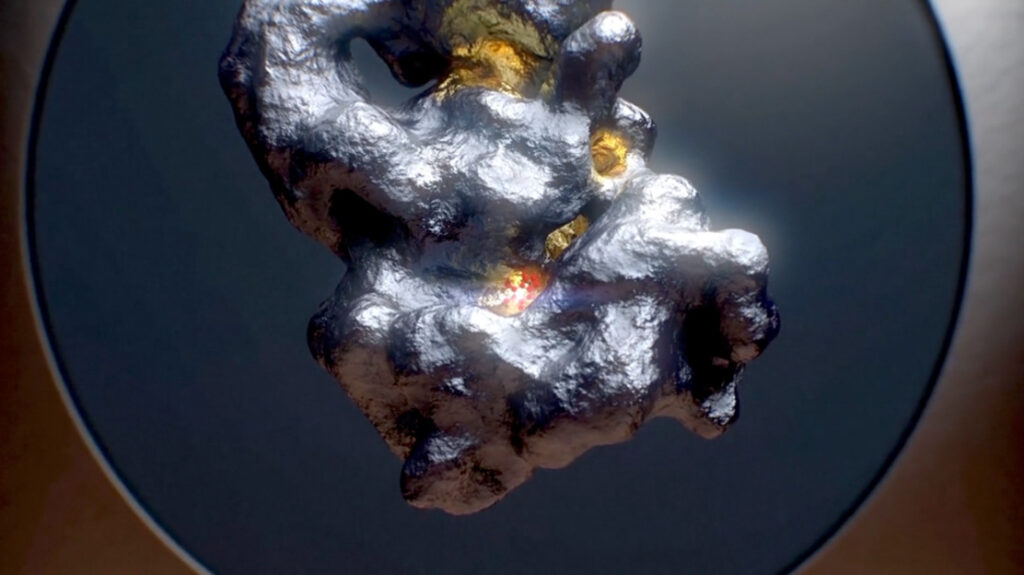

As a hit identification tool, ReSOLVE™ uniquely uses the hydrocophore™ to guide a virtual screen and efficiently identify potential hits. After hits are validated in the wet-lab, they drive additional screening and optimization of chemical matter, as demonstrated by our cGAS program. With ReSOLVE™, we are able to identify novel molecules for targets that have been inaccessible by small molecules to date, and quickly optimize these molecules towards a development candidate.